Over the past century, biomedical research has revealed that proteins are of central importance to disease and healthy human functioning. This case study, which was developed in partnership with St Vincent's Institute of Medical Research (SVI), describes how NHMRC-funded researchers at SVI revolutionised the sequencing of proteins.
As a consequence of their work, a vast collection of information on proteins is now available to researchers worldwide and is used daily to help develop more effective diagnostic techniques, therapeutic agents and devices.
A landscape format version of this case study is available as a PDF – see Download section below.
Origin
During the mid-1940s, at the University of Lund in Sweden, Dr Pehr Edman, a pioneer in the field of protein chemistry, became interested in how to quickly determine the detailed structure of protein molecules.
Protein molecules are themselves made up of chains of smaller molecules called amino acids. Prior to the 1950s, determining the sequence of amino acids that made up a single small protein was extremely difficult. Accomplishing this task was a 'chemical tour de force' that could require several years of work by an experienced and skilful chemist.1, 2
Edman discovered a series of three chemical reactions that would enable researchers to analyse any protein sequentially. He published this finding in 1950 in a paper that was read widely at the time and that remains frequently cited to this day. Edman's method (now known as Edman degradation) was instrumental in the rapid expansion internationally, in the 1950s and 1960s, of research in protein chemistry.3
In 1957, Edman moved to Melbourne to become the first Director of Research at the newly established School of Medical Research within St Vincent's Hospital (now St Vincent’s Institute of Medical Research (SVI)). He brought with him a focus on the structure and dynamics of proteins that continues to be a research focus for the Institute.
Grants and development
At SVI, Edman worked almost exclusively on further developing and improving the chemistry of his method.4 Other SVI research staff who supported and extended this work included Geoff Begg, Elizabeth Minasian, Agnes Henschen and Hugh Niall.
By 1960-61, the chemistry of the three-stage degradation reaction was well understood by the SVI team, but it was also clear that the manual application of this chemistry was slow and inefficient.
Moreover, the universal nature of the chemistry (it would work with any protein) and the repetitive nature of the sequencing process (the same three reactions were applied to each amino acid in turn) suggested to Edman and Begg independently that the whole process might be automated.
In addition, Edman realised that manually sequencing each of the many thousands of different types of proteins produced by human beings and other organisms was an impossible task. Automation was the only practical solution.5
Supported by grants from NHMRC, Edman's team worked to develop the world's first 'automatic protein amino-acid sequence analyser'6 which staff at SVI named the 'sequenator'.
A variety of technical problems had to be solved before automation of the many individual chemical reactions and other operations that had to occur within the machine was finally realised.
Critically, because of the flammable and corrosive liquids and vapours used as part of the degradation process, the sequenator had to be constructed using carefully chosen, chemically inert and highly resistant materials and with a focus on excluding or enclosing sparking electrical contacts.7,8
Results and translation
Edman and Begg published a full report of this work in 1967. The sequenator worked by adding solvents and reagents to the protein, which was spread out as a thin film on the inside wall of a spinning cylindrical glass cup. This process could produce an unbroken automatic sequencing of sixty amino acids at the rate of one per hour.
These results were considered 'stunning' at the time since the most rapid manual degradation previously possible occurred at the rate of one amino acid per day.9,10
The reported results were all the more impressive because, at that time, many laboratories found it too difficult to undertake even manual sequencing11 and because they could be achieved using a far smaller protein sample than had previously been required.12
As Edman reported to NHMRC in 1967, 'This instrument . . . simplifies enormously the problems of protein structure reducing years of work into weeks.'13

Commercialisation
At Edman's insistence, and with the agreement of SVI's board, the technology involved in the sequenator was not patented.14 'Home-made' automatic sequencers began to be constructed in a variety of other laboratories internationally, introducing modifications based on their own experiences.15
Encouraged by Edman and supported by an NHMRC fellowship, Niall went to work in the United States in 1967, first at the National Institutes of Health and then at Harvard Medical School.
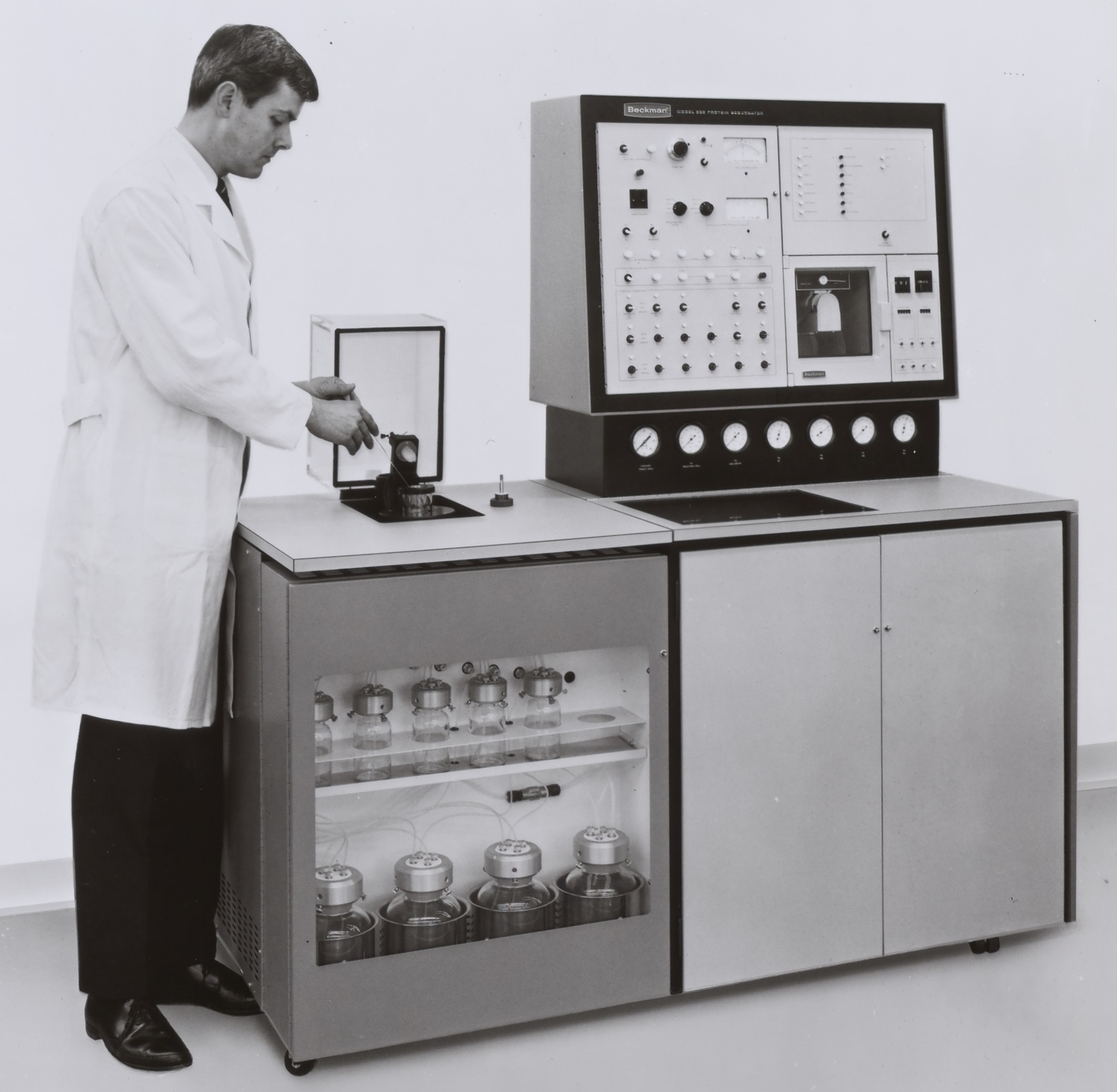
While in the US he collaborated with the Beckman instrument company on the development of a sequencer that was based on, and extended, Edman and Begg's original design.16

Outcomes and Impact
By 1973, there were over 100 instruments operating around the world based on the design of the sequenator. The most successful of these was the 890C, manufactured by Beckman Spinco under the guidance of Niall. At least four other companies in the USA also made such instruments, along with one Japanese company and another in Europe.17
Use of the Beckman sequencer and Edman chemistry dominated the field of structural biochemistry. Until the early 1990s, when mass spectrometry became available, Edman degradation was almost the only technique used for the direct determination of protein sequences.18
Automated protein sequencing (using various methods) has created vast amounts of biochemical data. Between 1949 and 1976, more than 80,000 amino acids had been sequenced by laboratories around the world.19 By 2017, this number had grown to at least 70 million.20
Protein sequencing has led to significant positive impacts on both knowledge and health. It has enabled characterisation of proteins for diagnostic and therapeutic use, and the understanding of protein mutations in genetic diseases such as phenylketonuria and thalassaemia.
Salcatonin (a hormone) was sequenced using the Beckman sequencer by Niall and now has widespread therapeutic use against osteoporosis, hypercalcemia and Paget's disease. Niall also sequenced enzymes, cytokines (immune system signalling proteins) and parathyroid hormone-related protein (which regulates bone and breast development, among its other functions).
Amino acid sequences obtained using Edman's technology enabled the cloning of many molecules used as drugs, including growth hormone and erythropoietin, a cytokine that stimulates red blood cell production in the bone marrow.21
Timeline
| When | Activity |
|---|---|
| 1950 | Edman publishes key paper on protein degradation |
| 1950-1957 | Edman undertakes experimental work to refine and demonstrate the detailed chemistry of his 3 stage method |
| 1957 | Edman moves to SVI |
| 1959-1961 | Grants (Edman) – Enzyme structure |
| 1962 | Grant (Edman) – Chemical structure of a short chain of amino acids |
| 1963 | Grant (Edman) – Sequencing chemistry |
| 1964, 1965 | Grant (Edman) – Sequencer development |
| 1966 | CJ Martin Travelling Fellowship (Niall) |
| 1967-1973 | Niall further develops automated sequencing in US |
| 1970, 1971, 1972 | Grants (Henschen) – sequencing of a very large protein (fibrinogen) |
| 1973 | Niall describes Beckman sequencer |
Proteins
Proteins are large, complex molecules that play many critical roles within the human body, including:
- antibodies (binding to foreign particles such as viruses and bacteria to help protect the body from infection)
- enzymes (carrying out most of the thousands of chemical reactions that take place in cells)
- messengers (in the form of hormones, transmitting signals to coordinate biological processes between different cells, tissues and organs)
- structural components (providing structure and support for cells and allowing the body to move)
- transport/storage (binding and carrying atoms and small molecules within cells and throughout the body).
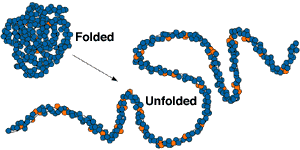
Each protein molecule is made up of a chain (or sequence) of amino acids. This chain folds to give the protein a unique three dimensional shape, which in turn determines its biological properties and behaviour.
The sequence of amino acids forming each protein is determined by the genetic code (DNA) of the organism that produces it. Human DNA may code for as many as 500,000 different proteins.22 The number of individual protein molecules in a human body is immense. For example, an individual human liver cell may contain about 10,000 different proteins, and a million molecules of each protein.23
Researcher profiles
Professor Pehr Edman
Pehr Victor Edman (1916-1977) received a Bachelor of Medicine and later a medical doctorate from the Karolinska Institute in Stockholm. After a fellowship at the Rockefeller Institute of Medical Research in the United States, he became an Associate Professor of Physiological Chemistry at the University of Lund. From 1957 to 1972 he was Director of Research at SVI and from 197 to 1977 he was Director of Protein Chemistry at the Max Planck Institute for Biochemistry in Munich. He was a fellow of the Royal Society, London, and the Australian Academy of Science, and was a recipient of a number of awards.
Geoff Begg
Geoff S Begg, a laboratory technician at SVI, received training at the Royal Melbourne Institute of Technology (RMIT) and was one of the first technical staff employed at SVI. Begg was an expert in practical chemistry, glassblowing, mechanical engineering and electronics.
Elizabeth Minasian
Elizabeth Minasian worked as a research assistant at SVI from 1962 to 1970 then at the Biochemistry Department of The University of Melbourne.
Professor Agnes Henschen
Agnes Henschen (d. 2011) studied and worked at the Karolinska Institute from 1952-1966, gaining a medical degree and doctorate. She left the Karolinska Institute to work at SVI in Melbourne, then in 1972 moved to the Max Planck Institute for Biochemistry. In 1989, she moved to the University of California, Irvine (UCI) where she remained until she retired in 2009. Henschen was instrumental in establishing UCI's first protein sequencing facility. She was a pioneer in the characterisation of the blood coagulation protein fibrinogen and was the first person to report its amino acid sequence.
Professor Hugh Niall
Hugh David Niall commenced studying medicine in 1954, then interrupted his studies to join SVI with Pehr Edman. After working in the United States, Niall returned to Australia in 1974 to the Howard Florey Institute. He moved to California in 1985 to Genetech (a biotechnology company) where he became Vice-President of Research Discovery. In 1995 he returned to Melbourne and became CEO of Biota Holdings, an Australian biotech company. He is now a Vice Chancellor's Professorial Fellow at Monash University.
References
This case study was developed in partnership with St Vincent’s Institute of Medical Research. The information and images from which impact case studies are produced may be obtained from a number of sources including our case study partner, NHMRC’s internal records and publicly available materials.
The following sources were consulted for this case study:
- Niall HD. Obituary Pehr Edman. Nature, 21 July 1977, vol 268
- Partridge SM, Blombäck B. Pehr Victor Edman, 14 April 1916 - 19 March 1977, Elected F.R.S. 1974. Biographical Memoirs of Fellows of the Royal Society. 1979;25:241-65. doi: 10.1098/rsbm.1979.0008. PMID: 11615794
- Moore S. Obituary Pehr Victor Edman. Trends in Biochemical Sciences, July 1977
- Morgan FJ. Pehr Victor Edman 1916-1977. Historical Records of Australian Science, 1990; 8(2)
- (4) Morgan
- (4) Morgan
- Niall HD. Automated Edman degradation: the protein sequenator. Comparative Study Methods Enzymol. 1973;27:942-1010. PMID: 4773306 DOI: 10.1016/s0076-6879(73)27039-8
- Edman P, Begg G. A protein sequenator. European Journal of Biochemistry, 1967;1(1):80-91. doi: 10.1007/978-3-662-25813-2_14. PMID: 6059350
- Geoff Begg, personal communication, 9 September 2021
- (2) Partridge
- (4) Morgan
- Doolittle, RF. An anecdotal account of the history of peptide stepwise degradation procedures. In M. Elzinga (ed.), Methods in Protein Sequence Analysis. The HUMANA Press Inc. 1982
- National Health and Medical Research Council. Thirtieth Annual Report for year 1967. Medical Research Endowment Act 1937. The Parliament of the Commonwealth of Australia 1968, Parliamentary Paper No. 199
- Sparrow L. Special feature on Edman. The legacy of Pehr Edman. Australian Biochemist, 2002, 33(2):16-21
- (7) Niall, (9) Doolittle, (14) Sparrow
- The University of Melbourne. Dr Hugh David Niall, Citation for the Award of Doctor of Medical Science (Honoris Causa). Faculty of Medicine, Dentistry and Health Sciences, Awards and Honours. 2012
- (7) Niall, (14) Sparrow
- (14) Sparrow
- (2) Partridge
- Pengfei T, Best RB. How Many Protein Sequences Fold to a Given Structure? A Coevolutionary Analysis. Biophys J . 2017;113(8):1719-1730. doi: 10.1016/j.bpj.2017.08.039. PMID: 29045866
- Van Der Weyden MB. Bench-to-bedside research in Australian research institutes: a snapshot. Med J Aust 2003; 179 (11): 603-610. doi: 10.5694/j.1326-5377.2003.tb05717.x
- Pray L. Eukaryotic genome complexity. Nature Education, 2008, 1(1):96
- Lodish H, Berk A, Zipursky SL, Matsudaira P, Baltimore D, Darnell J. Section 1.2. The molecules of life. Molecular Cell Biology, 4th Edition. New York: W. H. Freeman; 2000.
Partners
